Master’s thesis proposal – Microbial community composition in wastewater activated sludge at varying retention time
Background
Activated sludge (AS) is the most common method to treat wastewater and is globally the largest application of a biotechnological process. The activated sludge process relies on microorganisms to remove contaminants and the excess biomass that is formed is used for energy production as biogas. However, a large share of the microorganisms are kept in the system via the return sludge (see Figure). The retention time of the AS in the wastewater basins, referred to as solids retention time (SRT), needs to be higher than the time it takes for the microorganisms to multiply. Otherwise the microorganisms will be washed out from the wastewater treatment plant (WWTP). However, it can be desirable to have short SRT, because then a large share of the wastewater organic matter can be transformed to biogas, since less will be oxidized by the microorganisms to carbon dioxide (CO2) and lost in the atmosphere.
The SRT also determines the selection pressure on the microorganisms in the AS, where a short SRT is expected to select for fast growing microorganisms over slow growers. The microbial community composition in AS also depend on other factors that are set by the WWTP operator, such as aeration regimes, and on factors that are less controlled, such as the composition of microorganisms in the influent wastewater and seasonal changes (variations in temperature and rain which dilutes the wastewater). Despite >100 years of operation of AS plants, the knowledge about which microorganisms are present in AS at different conditions is still incomplete and the importance of the different factors is under debate.
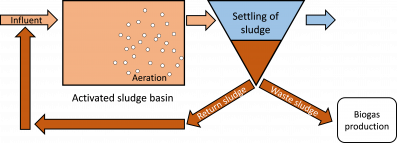
Figure. A simplified layout of an activated sludge wastewater treatment process.
Aim
This project focus on the effect of SRT on the evolution of the AS microbial communities. It will investigate sludge sampled from Sjölunda WWTP (Malmö, Sweden) at gradually increasing SRT in a parallel set-up consisting of several separate basins with separate AS. This provide a unique opportunity to compare full-scale processes and enables statistical evaluation.
The aim is to address the question whether the microbial community composition and dynamics is systematic in AS at decreasing SRT and provide data on which organisms thrive at which conditions. Established ecological theories will be tested as explanatory models of the obtained results.
Method
The microbial communities will be investigated by high throughput amplicon sequencing (MiSeq platform) of the 16S rRNA gene. This will provide data on the detailed community composition in the AS at a number of time points for several separate AS basins. By bioinformatics, communities will be compared and visualized using multivariate statistical tools. The workflow consists of labwork in molecular microbiology at Chalmers (DNA extraction, PCR amplification, gel-electrophoresis, DNA purification). Sequencing will be conducted at a Core Facility (outside the Chalmers lab). Bioinformatics will be performed using established pipelines in R and/or other environments.
When?
Spring 2017
Supervisor
Frank Persson, assistant professor, Water Environment Technology (WET), Civil and Environmental Engineering, Chalmers University of Technology, frank.persson@chalmers.se
David Gustavsson, research leader, Sweden Water Research, david.gustavsson@vasyd.se
